micro-sam
Segment Anything for microscopy — interactive and automatic segmentation built on Meta's Segment Anything model.
Overview
micro-sam adapts the Segment Anything foundation model to microscopy data with finetuned models. It cna run as a napari plugin for interactive segmentation of 2D, 3D, and time-series images, and also supports fine-tuning SAM on your own data for fully automatic instance segmentation. A recent GPU is strongly recommended.
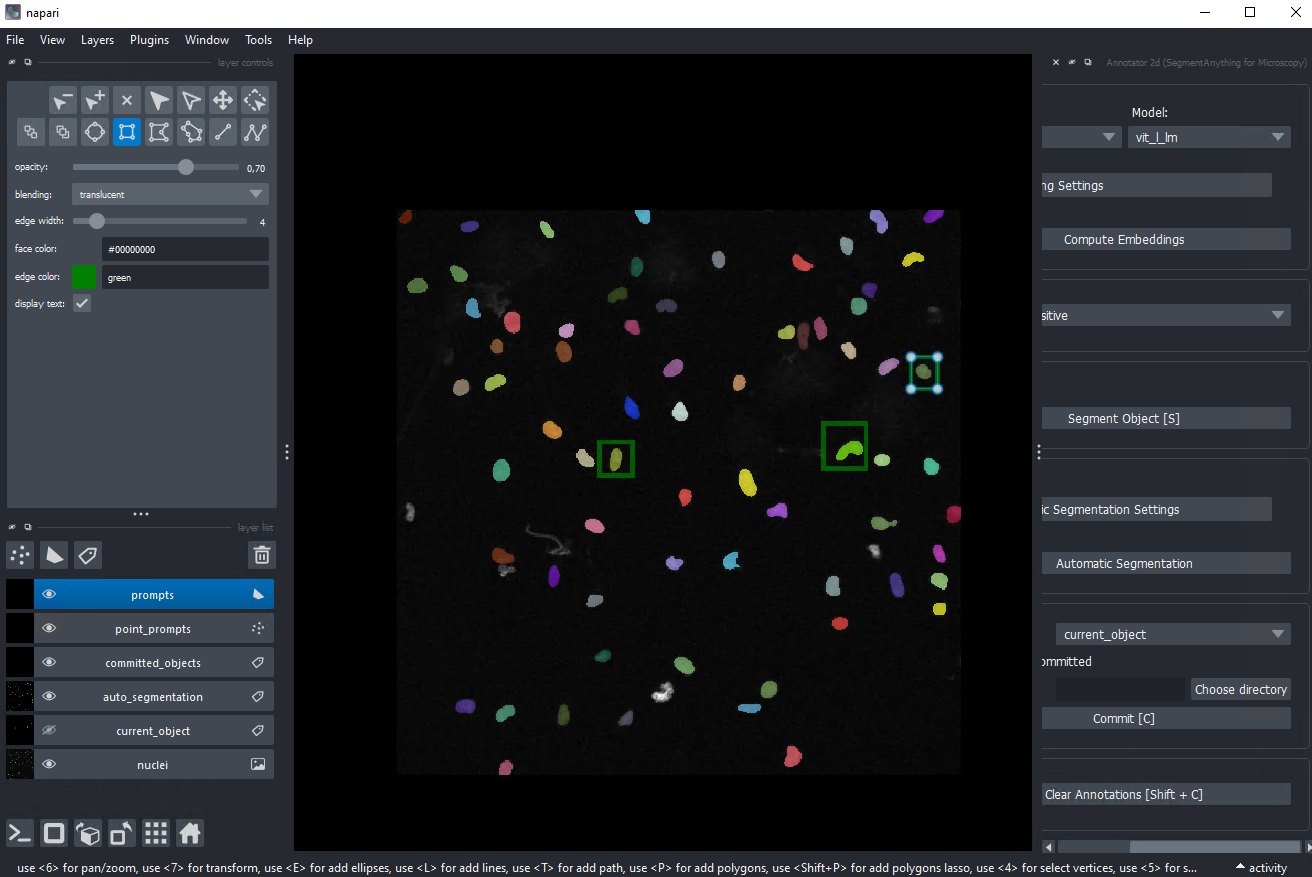
Key features
- Interactive point/box prompts for segmentation in napari
- Works on 2D, 3D volumes, and time-series
- Automatic instance segmentation after fine-tuning
- Pre-trained models specialised for light microscopy and electron microscopy
Installation
See Python environments for package manager setup (pixi or conda).
Recommended: use the ready-made pixi environment. The AI_tools_pixi repository ships a locked micro-sam environment with napari, trackastra, and napari-omero (CUDA 12.8):
git clone https://github.com/Leiden-Cell-Observatory/AI_tools_pixi
cd AI_tools_pixi/micro_sam
pixi install
pixi run napari
The micro-sam plugins appear under the napari Plugins menu.
Alternative: manual conda install.
conda install -c conda-forge micro_sam
napari
Tip
A CUDA-capable GPU makes interactive segmentation feel responsive; CPU-only is usable but slow, particularly on 3D data.
Official documentation
Learning resources
Written
- Annotation tools guide — the napari-based interactive workflow.
- Fine-tuning guide — train SAM on your own data for automatic segmentation.